
The same goes for the microfluidics devices that use beads. These beads are microscopic and each of them needs to be coated with a molecular barcode. For one sample, up to 3 million unique barcodes are required. In order to be able to offer single-cell sequencing in a commercially viable way, we needed to scale down the volume of the reagents used. This could only be done with sophisticated and expensive liquid handling robots. 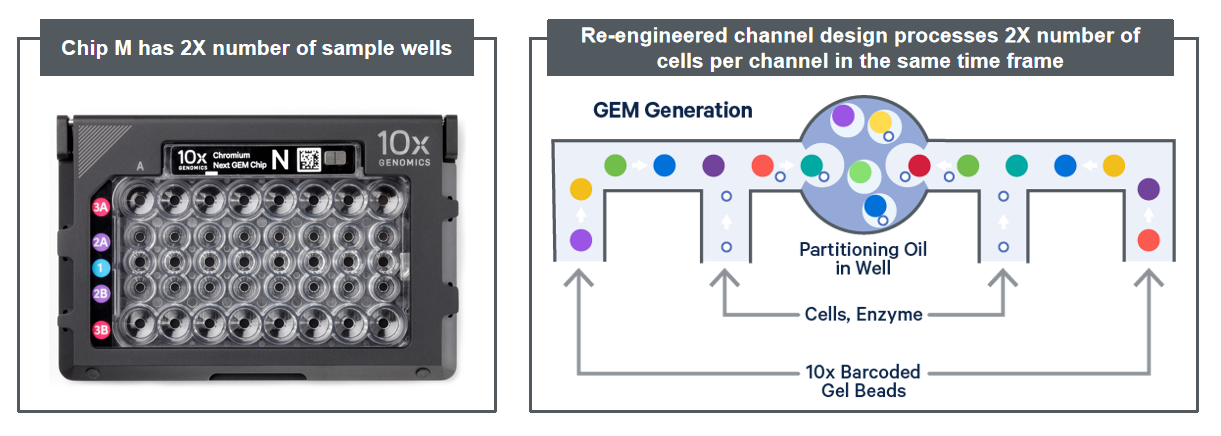
It’s a relatively new technology that only recently became available to a larger market thanks to a reduction in costs, as well as innovations in scaling up and standardization.īut, whatever platform or technology you choose, it is more sophisticated than conventional molecular biology technologies such as bulk RNA sequencing. The biological question you want to answer will determine which factor is most important. 
If you double the number of cells in your analysis, you will have to halve the number of reads per cell. This is one of the most important trade-offs to consider if you want to keep the sequencing costs stable. But to gain a thorough understanding of the transcriptome at the level of the individual cell, you need adequate sequencing depth to get a statistically significant dataset. Sequencing costs for single-cell sequencing are approximately 7 to 15 times higher than for traditional bulk analyses. That’s 7.5 to 15 times more than for a bulk RNA analysis! So, for example, a relatively simple single-cell sequencing project on the 10x Genomics platform, where you target 3,000 cells with a sequencing depth of 50,000 sequencing reads per cell, will need to generate 3,000 x 50,000 = 150 million reads. In contrast, the typical number of reads per cell needed for a single-cell sequencing project ranges from 50,000 to 150,000*. Just as with the reagents needed for library preparation, you will need many times more sequencing reads for a standard single-cell sequencing project than for a standard bulk RNA sequencing project.Ī typical bulk RNA sequencing analysis requires up to 20 million sequencing reads per sample. The cost of sequencing itself is often overlooked when considering the price of a single-cell sequencing project. Generally speaking, the reagent costs are 10-20 times higher for a standard single-cell RNA sequencing experiment than that of a bulk RNA experiment.įor single-cell sequencing, reagents costs are 10 to 20 times higher than for traditional bulk RNA sequencing. In contrast, when we analyze pooled cells in a single tube for a bulk RNA-seq experiment, we use much less reagent per cell. The costs of the reagentsįor a single-cell sequencing experiment, we process single cells in individual reactions, each of which requires its own set of reagents and this drastically increases the total amount of reagents needed. This post looks at the factors affecting the cost of your single-cell sequencing project, and how adapting the design of your experiment can keep costs down. Support Sample requirements, shipping, and more.Blog Thought-leadership, knowledge, and updates.Resource Library Webinars, our selection tool, and more.Case studies Detailed insights into past projects.Publications From our clients and project work.Getting started All the steps for starting your project.Data analysis Options beyond the included exploratory analysis.Our approach An inside look at SCD’s White-Glove approach.Research Areas Find out how single-cell sequencing is advancing disease understanding and drug development in your research area of interest.Applications Overview From fundamental biology to translational research and drug development, single-cell sequencing applications are revolutionizing science.Research & Development Our expert team drives innovation and versatility. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
